Stool testing has evolved significantly over the past decade. What was once limited to microscopic examination for ova and parasites now includes molecular detection of pathogens, quantification of commensal bacteria, assessment of digestive function, and measurement of intestinal inflammation.
The question is not whether stool testing can provide useful information — it can. The question is which tests provide actionable clinical data and which generate noise.
Types of Stool Testing
PCR-Based Comprehensive Stool Analysis
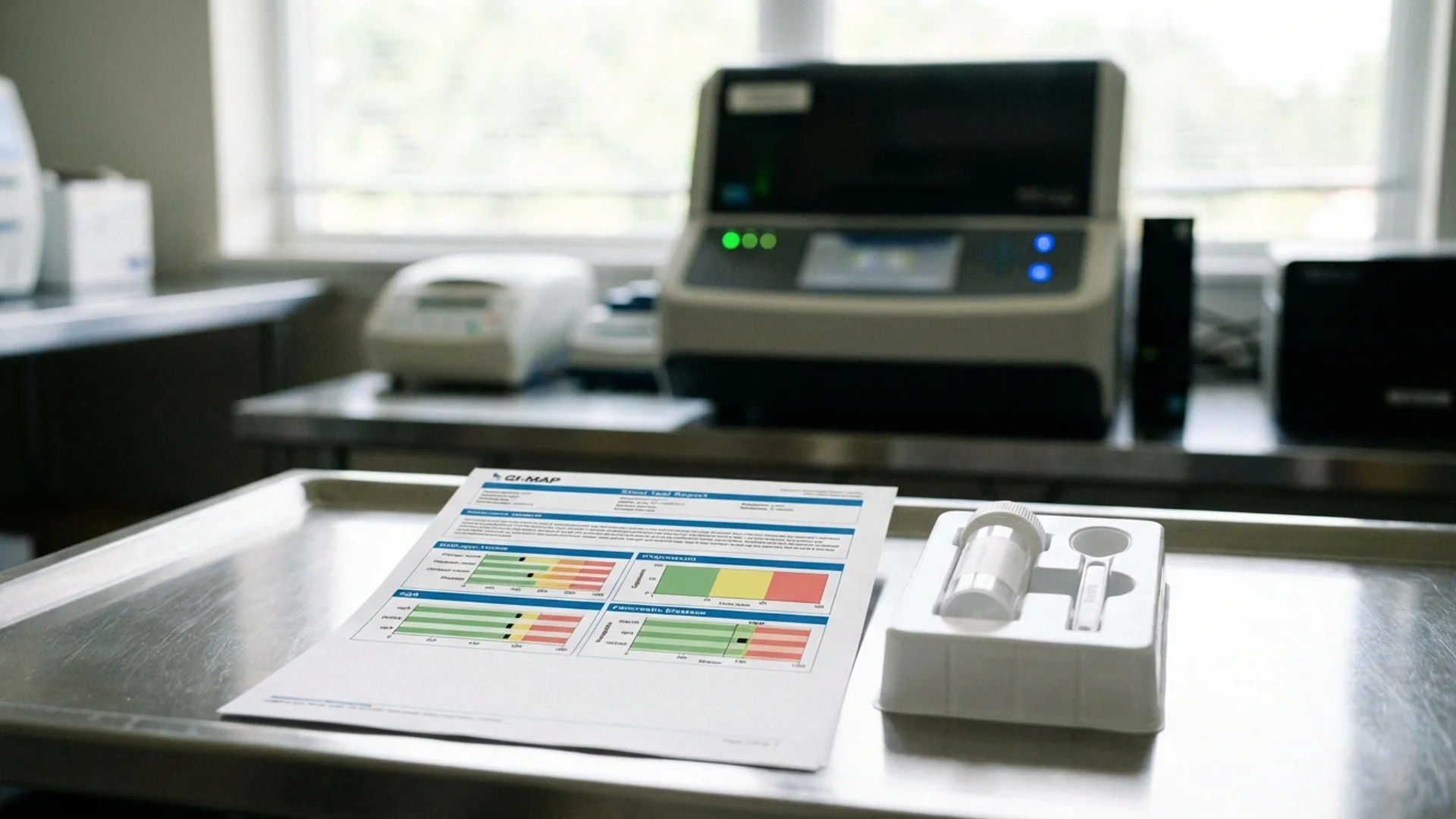
Tests like the GI-MAP (Diagnostic Solutions Laboratory) use quantitative PCR (qPCR) to detect and quantify specific organisms. This is the platform I use most frequently in clinical practice because it provides clinically actionable results.
What it measures:
- Bacterial pathogens — Clostridioides difficile, Helicobacter pylori (including virulence factors like CagA and VacA), Campylobacter, pathogenic E. coli strains, Vibrio, Yersinia
- Parasites — Giardia lamblia, Cryptosporidium, Entamoeba histolytica, Blastocystis hominis, Dientamoeba fragilis
- Viral pathogens — Norovirus, Rotavirus
- Opportunistic organisms — Candida species, Microsporidium, Pseudomonas, Staphylococcus, Streptococcus
- Commensal bacteria — quantification of beneficial species including Lactobacillus, Bifidobacterium, Bacteroides fragilis, Faecalibacterium prausnitzii, Akkermansia muciniphila, and others
- Inflammatory markers — calprotectin, lactoferrin, sIgA, anti-gliadin antibodies
- Digestive function markers — pancreatic elastase, steatocrit (fat in stool)
- Immune markers — secretory IgA, beta-glucuronidase
Advantages: High sensitivity for pathogen detection (superior to microscopy), quantitative results, comprehensive panel in a single sample. Evidence level for PCR-based pathogen detection: strong (validated against culture and other reference methods).
16S rRNA Sequencing
This technology sequences a conserved region of bacterial DNA to identify species composition. Companies like uBiome (no longer operating), Viome, and academic research laboratories use this approach.
What it measures: Relative abundance of bacterial species, microbial diversity indices (Shannon, Simpson), phylum-level composition (Firmicutes/Bacteroidetes ratio).
Limitations: Results are relative, not absolute (a decrease in one phylum appears as an increase in another, even if the actual amount has not changed). Clinical actionability is limited because the relationship between specific compositional patterns and clinical outcomes is still being characterized. Different laboratories use different methods, producing different results from the same sample.
My honest assessment: 16S sequencing is valuable for research but has limited clinical utility for individual patient management at this time. I do not routinely order it.
Shotgun Metagenomics
This sequences all DNA in the sample, providing species-level identification and functional gene information (what metabolic pathways the bacteria can perform).
Advantages: More detailed than 16S, provides functional information. Limitations: Expensive, data interpretation requires specialized expertise, and clinical translation frameworks are still being developed.
Key Markers and Their Interpretation
Calprotectin
Fecal calprotectin is released by neutrophils and is a specific marker of intestinal inflammation. It is one of the most validated stool markers available.
- Below 50 mcg/g — normal; inflammatory bowel disease is unlikely
- 50-200 mcg/g — indeterminate; may indicate mild inflammation, infection, or NSAID use
- Above 200 mcg/g — significant intestinal inflammation; further evaluation (typically colonoscopy) is warranted
Evidence level: strong (multiple systematic reviews and meta-analyses confirm its diagnostic accuracy for differentiating IBD from IBS). Sensitivity for IBD is approximately 93%; specificity approximately 96%.
Secretory IgA (sIgA)
sIgA reflects mucosal immune function. Low sIgA indicates impaired mucosal defense and is associated with increased susceptibility to gut infections and food sensitivities. Elevated sIgA may indicate acute infection or immune activation in the gut.
Pancreatic Elastase
Elastase is produced by the pancreas and remains stable during intestinal transit. Low pancreatic elastase (below 200 mcg/g) indicates exocrine pancreatic insufficiency — the pancreas is not producing sufficient digestive enzymes. This finding changes management directly: pancreatic enzyme replacement therapy addresses the deficiency.
Helicobacter pylori
The GI-MAP detects H. pylori DNA along with virulence factors. CagA-positive and VacA-positive strains are associated with higher risk of gastric ulceration and gastric cancer. Detecting these virulence factors influences treatment decisions.
Beta-Glucuronidase
This bacterial enzyme deconjugates compounds (including hormones) that the liver has conjugated for excretion. Elevated beta-glucuronidase leads to reabsorption of estrogen and toxins that should have been excreted — contributing to estrogen dominance and impaired detoxification. This marker connects gut health to hormonal balance in a clinically meaningful way.
Clinical Interpretation
The art of stool test interpretation lies not in reading individual markers but in synthesizing the overall picture.
Example 1: A patient with chronic fatigue and bloating. Results show elevated Candida albicans, low Lactobacillus and Bifidobacterium, low pancreatic elastase, and low sIgA. The interpretation: fungal overgrowth in the context of depleted beneficial flora, insufficient digestive enzyme production, and impaired mucosal immunity. Treatment: antifungal therapy, digestive enzyme supplementation, probiotic restoration, and mucosal immune support.
Example 2: A patient with alternating diarrhea and constipation. Results show elevated calprotectin (180 mcg/g), positive for Blastocystis hominis, low Faecalibacterium prausnitzii, and elevated sIgA. The interpretation: subclinical intestinal inflammation, possibly driven by Blastocystis infection, with depleted butyrate-producing bacteria. Treatment: address the parasitic infection, support anti-inflammatory gut flora, investigate further if calprotectin does not normalize.
Practical Recommendations
- Start with a PCR-based comprehensive stool analysis if you are investigating gut dysfunction. It provides the most clinically actionable information per test.
- Use calprotectin as a standalone test when the primary question is “is there significant intestinal inflammation?” — particularly when differentiating IBS from IBD.
- Retest at 8-12 weeks after treatment to confirm that interventions have produced the expected changes. Treating without follow-up testing means flying blind.
- Do not over-interpret sequencing data. If a sequencing report tells you that your Firmicutes/Bacteroidetes ratio is abnormal, that information alone does not tell me what to treat.
- Context matters. A positive finding for Blastocystis hominis in an asymptomatic patient may not require treatment. The same finding in a patient with chronic GI symptoms and elevated calprotectin likely does.
Disclaimer: This article is provided for educational purposes and reflects one physician’s clinical perspective. It is not a substitute for individualized medical care. Stool test results should be interpreted by a qualified physician in the context of clinical symptoms.




